
|

|
|
| Home | Live Data | Instruments | CHRNS | Proposals |
Introduction
A new bioengineering research partnership, Cold Neutrons for Biology and Technology (CNBT), consisting of investigators from five universities: University of California Irvine, University of Pennsylvania, Rice University, Duke University and Carnegie-Mellon University and NIST has recently been funded by the NIH. The CNBT partnership is committed to the development of advanced neutron scattering instrumentation devoted to basic and applied studies of membranes and macromolecules in membranes, and to membrane-based technologies that include studies of protein complexes with relevance to bioengineering. A next-generation advanced neutron diffractometer/ reflectometer, fully dedicated to biological membrane studies, has already been designed and will be built at the NIST Center for Neutron Research (NCNR) during the next two years. In addition, 10% of a state-of-the-art small-angle neutron scattering (SANS) spectrometer, currently operating at the NCNR, will be dedicated to studies of biological membrane systems. New sample environments, optimized for membrane research, will be designed and built under the partnership.
Biological structure studies will be augmented by advanced molecular dynamics methods in order to produce three-dimensional structural models from the neutron diffraction or SANS data. A high-performance computer system will be put in place at the NCNR for this purpose. The CNBT instrumentation and computing facilities will provide new research capabilities not currently available in the United States, leading to important advances in the structural studies of membranes and membrane proteins. These advances are expected to have an impact on a number of bioengineering applications including pharmaceutical design and screening, tissue engineering and biosensors.
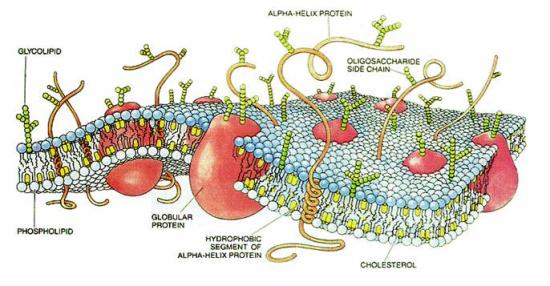
Cartoon of a typical cell membrane. Proteins are embedded in a matrix of phospholipid molecules. Several other membrane constituents such as glycolipids and cholesterol are also shown. Due to the complex nature of cell membranes, experiments to understand the structure and function of membrane proteins are performed on simpler biomimetic systems.
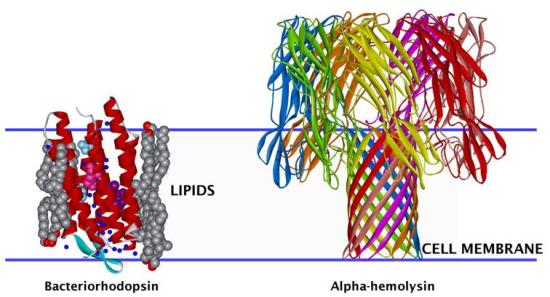
3-D structural representation of membrane proteins of two types: alpha-helical bacteriorhodopsin, a light-driven ion pump, and beta-barrel alpha-hemolysin, a bacterial toxin. Their disposition relative to the cell membrane is indicated by the blue lines. The structures of both proteins were determined by high-resolution x-ray crystallography. However, in general, it is difficult to obtain 3-D structures of membrane proteins in this manner due to the fluid nature of membranes and consequent lack of crystalline order in the membrane plane. Thus, a combination of neutron and x-ray diffraction or reflectivity data with molecular dynamics simulations must be used to arrive at 3-D models for membrane protein structure.
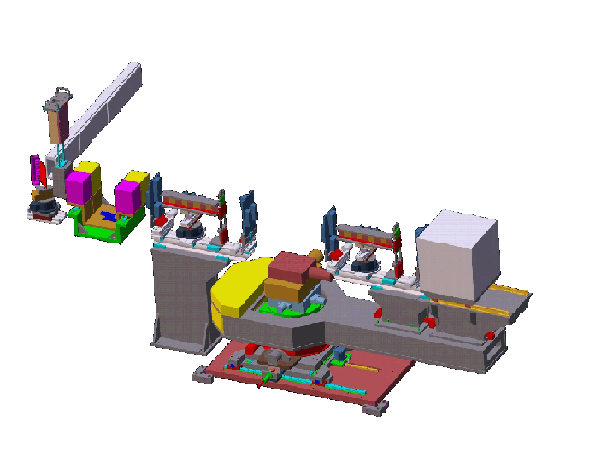

Design drawing of the CNBT advanced neutron diffractometer/reflectometer (AND/R). The instrument design follows that of the NCNR’s current state-of-the art NG1 reflectometer, but is is optimized for studies of biomembranes and biomembrane components.
Reflectivity (left) and scattering length density (SLD) profiles (right) for thiahexaethylene oxide alkane/d54-DMPC hybrid bilayer membranes (HBMs) in the absence (black) and presence (blue) of melittin, a small peptide toxin. The neutron results show that no water penetrates into the highly-ordered alkane layer. While the main effect of melittin appears to be dehydration of the DMPC head group layer, melittin clearly has an effect on the alkane/DMPC interface as well. The HBMs were deposited on silicon substrates that had been coated with 50Ĺ of gold on a 10Ĺ chromium layer. The gold and chromium layers are not shown in the SLD profiles for clarity.
Diffraction peaks from an ordered multilayer sample of DOPC deposited on a silicon substrate and measured at 44% relative humidity.
CNBT Projects:
a) Construct, optimize and operate a unique advanced neutron diffractometer/ reflectometer (AND/R) for structural studies of oriented monolayer and multil amellar membrane systems.
b) Advance the development of neutron diffraction methods for the study of supramolecular peptide-protein assemblies in membranes using SANS, AND/R and triple-axis spectrometers.
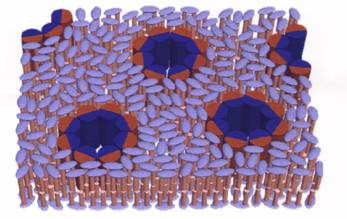
c) As a means of optimizing and expanding the capabilities of the AND/R, enhance, refine and expand methods for the determination of the structure of peptide-fluid-bilayer complexes by joint refinement of neutron, X-ray and molecular dynamics data.
d) Develop and refine the use of specular neutron interferometry, off-specular neutron scattering and molecular dynamics simulations methods for determining the secondary and tertiary structures of small proteins vectorially oriented in single-membrane systems.
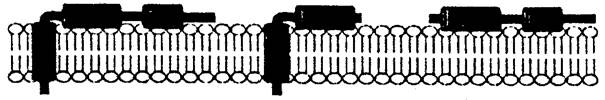
e) Enhance the use of neutron diffraction, reflectivity and interferometry as tools for the development of hybrid bilayer membranes (HBMs) as biomimetic surfaces for applications such as sensing and pharmaceutical screening.
Associated Collaborative Arrangements Resource for Solid-State NMR of Proteins at University of California San Diego, S. Opella, PI NMR Studies of Lipid Bilayers at the National Institute of Alcohol Abuse and Alcoholism at the NIH, K. Gawrisch, PI NSF Materials Research Science and Engineering Program at the University of Pennsylvania, M. Klein, PI National Stable Isotope Resource at Los Alamos National Laboratory
Last modified 31-August-2016 by website owner: NCNR (attn: Brian Kirby)